Bold new microscopies for the brain
New technologies are changing how scientists study the brain’s fine structure

McGovern researchers create unexpected new approaches to microscopy that are changing the way scientists look at the brain.
Ask McGovern Investigator Ed Boyden about his ten-year plan and you’ll get an immediate and straight-faced answer: “We would like to understand the brain.”
He means it. Boyden intends to map all of the cells in a brain, all of their connections, and even all of the molecules that form those connections and determine their strengths. He also plans to study how information flows through the brain and to use this to generate a working model. “I’d love to be able to load a map of an entire brain into a computer and see if we can simulate the brain,” he says.
Boyden likens the process to reverse-engineering a computer by opening it up and looking inside. The analogy, though not perfect, provides a sense of the enormity of the task ahead. As complicated as computers are, brains are far more complex, and they are also much harder to visualize, given the need to see features at multiple scales. For example, signals travel from cell to cell through synaptic connections that are measured in nanometers, but the signals are then propagated along nerve fibers that may span several centimeters—a difference of more than a million-fold. Modern microscopes make it possible to study features at one scale or the other, but not both together. Similarly, there are methods for visualizing electrical activity in single neurons or in whole brains, but there is no way to see both at once. So Boyden is building his own tools, and in the process is pushing the limits of imagination. “Our group is often trying to do the opposite of what other people do,” Boyden says.
Boyden’s new methods are part of a broader push to understand the brain’s connectivity, an objective that gained impetus two years ago with the President’s BRAIN Initiative, and with allied efforts such as the NIH-funded Human Connectome Project. Hundreds of researchers have already downloaded Boyden’s recently published protocols, including colleagues at the McGovern Institute who are using them to advance their studies of brain function and disease.
Just add water
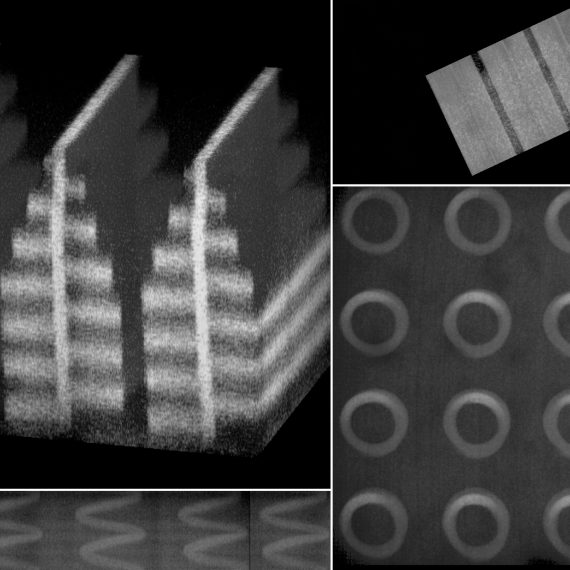
Under the microscope, the brain section prepared by Jill Crittenden looks like a tight bundle of threads. The nerve fibers are from a mouse brain, from a region known to degenerate in humans with Parkinson’s disease. The loss of the tiny synaptic connections between these fibers may be the earliest signs of degeneration, so Crittenden, a research scientist who has been studying this disease for several years in the lab of McGovern Investigator Ann Graybiel, wants to be able to see them.
But she can’t. They are far too small— smaller than a wavelength of light, meaning they are beyond the limit for optical microscopy. To bring these structures into view, one of Boyden’s technologies, called expansion microscopy (ExM), simply makes the specimen bigger, allowing it to be viewed on a conventional laboratory microscope.
The idea is at once obvious and fantastical. “Expansion microscopy is the kind of thing scientists daydream about,” says Paul Tillberg, a graduate student in Boyden’s lab. “You either shrink the scientist or expand the specimen.”
Leaving Crittenden’s sample in place, Tillberg adds water. Minutes later, the tissue has expanded and become transparent, a ghostly and larger version of its former self.
Crittenden takes another look through the scope. “It’s like someone has loosened up all the fibers. I can see each one independently, and see them interconnecting,” she says. “ExM will add a lot of power to the tools we’ve developed for visualizing the connections we think are degenerating.”
It took Tillberg and his fellow graduate student Fei Chen several months of brainstorming to find a plausible way to make ExM a reality. They had found inspiration in the work of MIT physicist Toyoichi Tanaka, who in the 1970s had studied smart gels, polymers that rapidly expand in response to a change in environment. One familiar example is the absorbent material in baby diapers, and Boyden’s team turned to this substance for the expansion technique.
The process they devised involves several steps. The tissue is first labeled using fluorescent antibodies that bind to molecules of interest, and then it is impregnated with the gel-forming material. Once the gel has set, the fluorescent markers are anchored to the gel, and the original tissue sample is digested, allowing the gel to stretch evenly in all directions.
When water is added, the gel expands and the fluorescent markers spread out like a picture on a balloon. Remarkably, the 3D shapes of even the finest structures are faithfully preserved during the expansion, making it possible to see them using a conventional microscope. By labeling molecules with different colors, the researchers can even distinguish pre-synaptic from post-synaptic structures. Boyden plans eventually to use hundreds, possibly thousands, of colors, and to increase the expansion factor to 10 times original size, equivalent to a 1000-fold increase in volume.
ExM is not the only way to see fine structures such as synapses; they can also be visualized by electron microcopy, or by recently-developed ‘super-resolution’ optical methods that garnered a 2014 Nobel Prize. These techniques, however, require expensive equipment, and the images are very time-consuming to produce.
“With ExM, because the sample is physically bigger, you can scan it very quickly using just a regular microscope,” says Boyden.
Boyden is already talking to other leading researchers in the field, including Kwanghun Chung at MIT and George Church at Harvard, about ways to further enhance the ExM method. Within the McGovern Institute, among those who expect to benefit from these advances is Guoping Feng, who is developing mouse models of autism, schizophrenia and other disorders by introducing some of the same genetic changes seen in humans with these disorders. Many of the genes associated with autism and schizophrenia play a role in the formation of synapses, but even with the mouse models at his disposal, Feng isn’t sure what goes wrong with them because they are so hard to see. “If we can make parts of the brain bigger, we might be able to see how the assembly of this synaptic machinery changes in different disorders,” he says.
3D Movies Without Special Glasses
Another challenge facing Feng and many other researchers is that many brain functions, and many brain diseases, are not confined to one area, but are widely distributed across the brain. Trying to understand these processes by looking through a small microscopic window has been compared to watching a soccer game by observing just a single square foot of the playing field.
No current technology can capture millisecond-by-millisecond electrical events across the entire living brain, so Boyden and collaborators in Vienna, Austria, decided to develop one. They turned to a method called light field microscopy (LFM) as a way to capture 3D movies of an animal’s thoughts as they flash through the entire nervous system.
The idea is mind-boggling to imagine, but the hardware is quite simple. The instrument records images in depth the same way humans do, using multiple ‘eyes’ to send slightly offset 2D images to a computer that can reconstruct a 3D image of the world. (The idea had been developed in the 1990s by Boyden’s MIT colleague Ted Adelson, and a similar method was used to create Google Street View.) Boyden and his collaborators started with a microscope of standard design, attached a video camera, and inserted between them a six-by-six array of miniature lenses, designed in Austria, that projects a grid of offset images into the camera and the computer.
The rest is math. “We take the multiple, superimposed flat images projected through the lens array and combine them into a volume,” says Young-Gyu Yoon, a graduate student in the Boyden lab who designed and wrote the software.
Another graduate student, Nikita Pak, used the new method to measure neural activity in C. elegans, a tiny worm whose entire nervous system consists of just 302 neurons. By using a worm that had been genetically engineered so that its neurons light up when they become electrically active, Pak was able to make 3D movies of the activity in the entire nervous system. “The setup is just so simple,” he says. “Every time I use it, I think it’s cool.”
The team then tested their method on a larger brain, that of the larval zebra fish. They presented the larvae with a noxious odor, and found that it triggered activity in around 5000 neurons, over a period of about three minutes. Even with this relatively simple example, activity is distributed widely throughout the brain, and would be difficult to detect with previous techniques. Boyden is now working towards recording activity over much longer timespans, and he also envisions scaling it up to image the much more complex brains of mammals.
He hopes to start with the smallest known mammal, the Etruscan shrew. This animal resembles a mouse, but it is ten times smaller, no bigger than a thimble. Its brain is also much smaller, with only a few million neurons, compared to 100 million in a mouse.
Whole brain imaging in this tiny creature could provide an unprecedented view of mammalian brain activity, including its disruption in disease states. Feng cites sensory overload in autism as an example. “If we can see how sensory activity spreads through the brain, we can start to understand how overload starts and how it spills over to other brain areas,” he says.
Visions of Convergence
While Boyden’s microscopy technologies are providing his colleagues with new ways to study brain disorders, Boyden himself hopes to use them to understand the brain as a whole. He plans to use ExM to map connections and identify which molecules are where; 3D whole-brain imaging to trace brain activity as it unfolds in real time, and optogenetics techniques to stimulate the brain and directly record the resulting activity. By combining all three tools together, he hopes to pin stimuli and activity to the molecules and connections on the map and then use that to build a computational model that simulates brain activity.
The plan is grandiose, and the tools aren’t all ready yet, but to make the scheme plausible in the proposed timeframe, Boyden is adhering to a few principles. His methods are fast, capturing information-dense images rapidly rather than scanning over days, and inclusive, imaging whole brains rather than chunks that need to be assembled. They are also accessible, so researchers don’t need to spend large sums to acquire specialized equipment or expertise in-house.
The challenges ahead might appear insurmountable at times, but Boyden is undeterred. He moves forward, his mind open to even the most far-fetched ideas, because they just might work.