Researchers advance CRISPR-based tool for diagnosing disease
With SHERLOCK, a strip of paper can now indicate presence of pathogens, tumor DNA, or any genetic signature of interest.
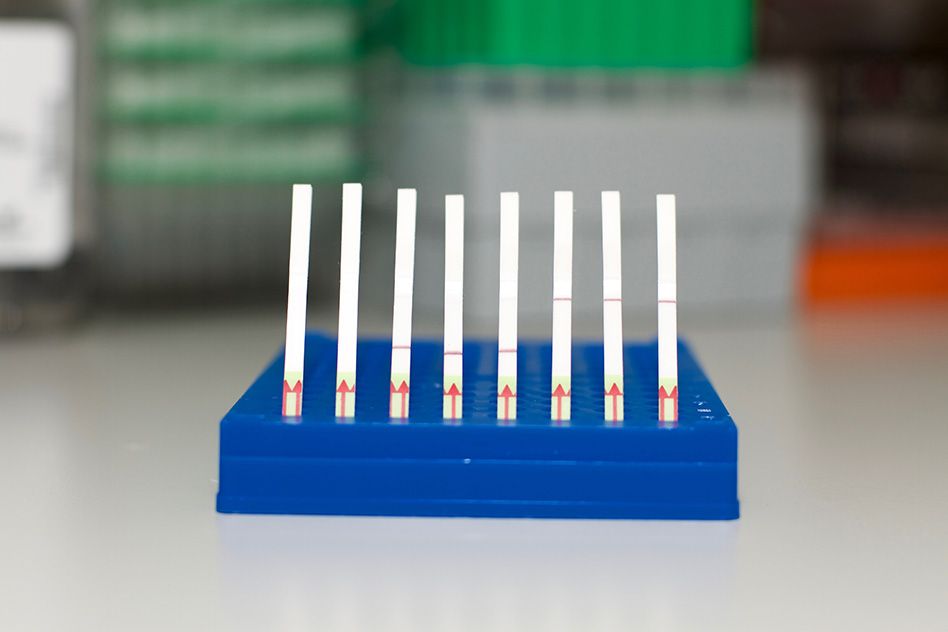

The team that first unveiled the rapid, inexpensive, highly sensitive CRISPR-based diagnostic tool called SHERLOCK has greatly enhanced the tool’s power, and has developed a miniature paper test that allows results to be seen with the naked eye — without the need for expensive equipment.
The SHERLOCK team developed a simple paper strip to display test results for a single genetic signature, borrowing from the visual cues common in pregnancy tests. After dipping the paper strip into a processed sample, a line appears, indicating whether the target molecule was detected or not.
This new feature helps pave the way for field use, such as during an outbreak. The team has also increased the sensitivity of SHERLOCK and added the capacity to accurately quantify the amount of target in a sample and test for multiple targets at once. All together, these advancements accelerate SHERLOCK’s ability to quickly and precisely detect genetic signatures — including pathogens and tumor DNA — in samples.
Described today in Science, the innovations build on the team’s earlier version of SHERLOCK (shorthand for Specific High Sensitivity Reporter unLOCKing) and add to a growing field of research that harnesses CRISPR systems for uses beyond gene editing. The work, led by researchers from the Broad Institute of MIT and Harvard and from MIT, has the potential for a transformative effect on research and global public health.
“SHERLOCK provides an inexpensive, easy-to-use, and sensitive diagnostic method for detecting nucleic acid material — and that can mean a virus, tumor DNA, and many other targets,” said senior author Feng Zhang, a core institute member of the Broad Institute, an investigator at the McGovern Institute, and the James and Patricia Poitras ’63 Professor in Neuroscience and associate professor in the departments of Brain and Cognitive Sciences and Biological Engineering at MIT. “The SHERLOCK improvements now give us even more diagnostic information and put us closer to a tool that can be deployed in real-world applications.”
The researchers previously showcased SHERLOCK’s utility for a range of applications. In the new study, the team uses SHERLOCK to detect cell-free tumor DNA in blood samples from lung cancer patients and to detect synthetic Zika and Dengue virus simultaneously, in addition to other demonstrations.
Clear results on a paper strip
“The new paper readout for SHERLOCK lets you see whether your target was present in the sample, without instrumentation,” said co-first author Jonathan Gootenberg, a Harvard graduate student in Zhang’s lab as well as the lab of Broad core institute member Aviv Regev. “This moves us much closer to a field-ready diagnostic.”
The team envisions a wide range of uses for SHERLOCK, thanks to its versatility in nucleic acid target detection. “The technology demonstrates potential for many health care applications, including diagnosing infections in patients and detecting mutations that confer drug resistance or cause cancer, but it can also be used for industrial and agricultural applications where monitoring steps along the supply chain can reduce waste and improve safety,” added Zhang.
At the core of SHERLOCK’s success is a CRISPR-associated protein called Cas13, which can be programmed to bind to a specific piece of RNA. Cas13’s target can be any genetic sequence, including viral genomes, genes that confer antibiotic resistance in bacteria, or mutations that cause cancer. In certain circumstances, once Cas13 locates and cuts its specified target, the enzyme goes into overdrive, indiscriminately cutting other RNA nearby. To create SHERLOCK, the team harnessed this “off-target” activity and turned it to their advantage, engineering the system to be compatible with both DNA and RNA.
SHERLOCK’s diagnostic potential relies on additional strands of synthetic RNA that are used to create a signal after being cleaved. Cas13 will chop up this RNA after it hits its original target, releasing the signaling molecule, which results in a readout that indicates the presence or absence of the target.
Multiple targets and increased sensitivity
The SHERLOCK platform can now be adapted to test for multiple targets. SHERLOCK initially could only detect one nucleic acid sequence at a time, but now one analysis can give fluorescent signals for up to four different targets at once — meaning less sample is required to run through diagnostic panels. For example, the new version of SHERLOCK can determine in a single reaction whether a sample contains Zika or dengue virus particles, which both cause similar symptoms in patients. The platform uses Cas13 and Cas12a (previously known as Cpf1) enzymes from different species of bacteria to generate the additional signals.
SHERLOCK’s second iteration also uses an additional CRISPR-associated enzyme to amplify its detection signal, making the tool more sensitive than its predecessor. “With the original SHERLOCK, we were detecting a single molecule in a microliter, but now we can achieve 100-fold greater sensitivity,” explained co-first author Omar Abudayyeh, an MIT graduate student in Zhang’s lab at Broad. “That’s especially important for applications like detecting cell-free tumor DNA in blood samples, where the concentration of your target might be extremely low. This next generation of features help make SHERLOCK a more precise system.”
The authors have made their reagents available to the academic community through Addgene and their software tools can be accessed via the Zhang lab website and GitHub.
This study was supported in part by the National Institutes of Health and the Defense Threat Reduction Agency.