McGovern neuroscientists develop a new model for autism
By introducing a gene variant associated with autism into monkeys, researchers hope to study treatment options for severe neurodevelopmental disorders.
Using the genome-editing system CRISPR, researchers at MIT and in China have engineered macaque monkeys to express a gene mutation linked to autism and other neurodevelopmental disorders in humans. These monkeys show some behavioral traits and brain connectivity patterns similar to those seen in humans with these conditions.
Mouse studies of autism and other neurodevelopmental disorders have yielded drug candidates that have been tested in clinical trials, but none of them have succeeded. Many pharmaceutical companies have given up on testing such drugs because of the poor track record so far.
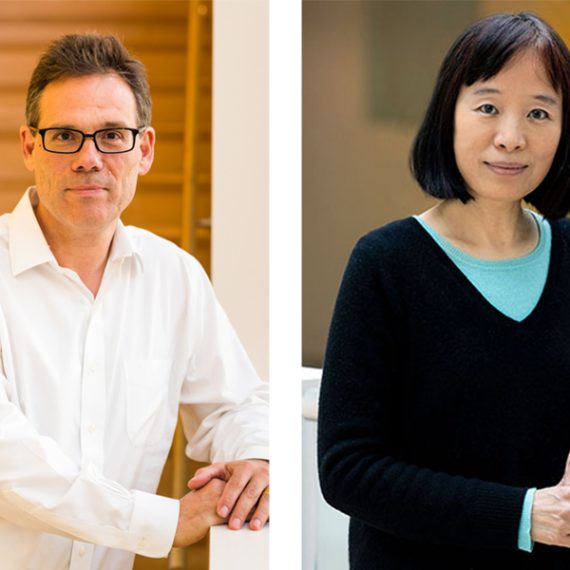
The new type of model, however, could help scientists to develop better treatment options for some neurodevelopmental disorders, says Guoping Feng, who is the James W. and Patricia Poitras Professor of Neuroscience, a member of MIT’s McGovern Institute for Brain Research, and one of the senior authors of the study.
“Our goal is to generate a model to help us better understand the neural biological mechanism of autism, and ultimately to discover treatment options that will be much more translatable to humans,” says Feng, who is also an institute member of the Broad Institute of MIT and Harvard and a senior scientist in the Broad’s Stanley Center for Psychiatric Research.
“We urgently need new treatment options for autism spectrum disorder, and treatments developed in mice have so far been disappointing. While the mouse research remains very important, we believe that primate genetic models will help us to develop better medicines and possibly even gene therapies for some severe forms of autism,” says Robert Desimone, the director of MIT’s McGovern Institute for Brain Research, the Doris and Don Berkey Professor of Neuroscience, and an author of the paper.
Huihui Zhou of the Shenzhen Institutes of Advanced Technology, Andy Peng Xiang of Sun Yat-Sen University, and Shihua Yang of South China Agricultural University are also senior authors of the study, which appears in the June 12 online edition of Nature. The paper’s lead authors are former MIT postdoc Yang Zhou, MIT research scientist Jitendra Sharma, Broad Institute group leader Rogier Landman, and Qiong Ke of Sun Yat-Sen University. The research team also includes Mriganka Sur, the Paul and Lilah E. Newton Professor in the Department of Brain and Cognitive Sciences and a member of MIT’s Picower Institute for Learning and Memory.
Gene variants
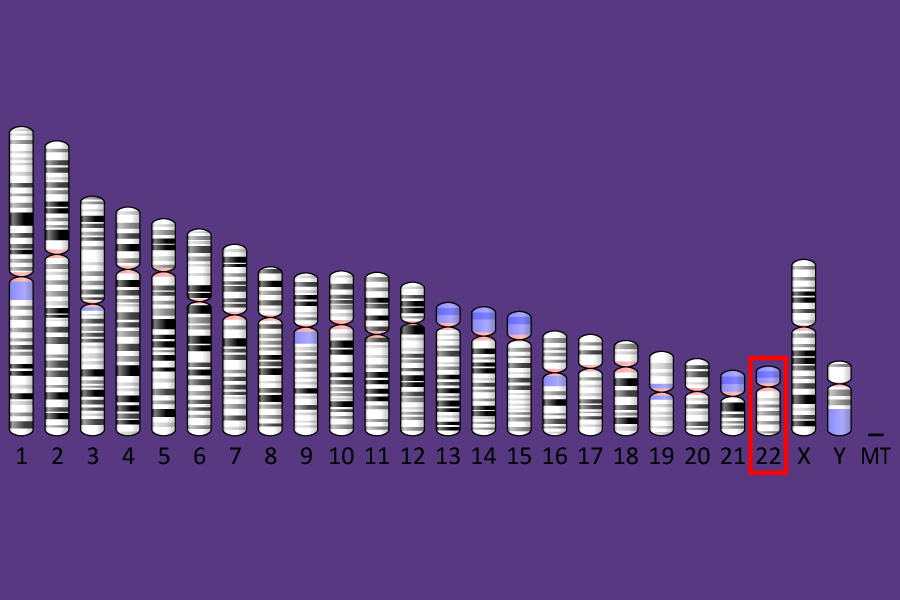
Scientists have identified hundreds of genetic variants associated with autism spectrum disorder, many of which individually confer only a small degree of risk. In this study, the researchers focused on one gene with a strong association, known as SHANK3. In addition to its link with autism, mutations or deletions of SHANK3 can also cause a related rare disorder called Phelan-McDermid Syndrome, whose most common characteristics include intellectual disability, impaired speech and sleep, and repetitive behaviors. The majority of these individuals are also diagnosed with autism spectrum disorder, as many of the symptoms overlap.
The protein encoded by SHANK3 is found in synapses — the junctions between brain cells that allow them to communicate with each other. It is particularly active in a part of the brain called the striatum, which is involved in motor planning, motivation, and habitual behavior. Feng and his colleagues have previously studied mice with Shank3 mutations and found that they show some of the traits associated with autism, including avoidance of social interaction and obsessive, repetitive behavior.
Although mouse studies can provide a great deal of information on the molecular underpinnings of disease, there are drawbacks to using them to study neurodevelopmental disorders, Feng says. In particular, mice lack the highly developed prefrontal cortex that is the seat of many uniquely primate traits, such as making decisions, sustaining focused attention, and interpreting social cues, which are often affected by brain disorders.
The recent development of the CRISPR genome-editing technique offered a way to engineer gene variants into macaque monkeys, which has previously been very difficult to do. CRISPR consists of a DNA-cutting enzyme called Cas9 and a short RNA sequence that guides the enzyme to a specific area of the genome. It can be used to disrupt genes or to introduce new genetic sequences at a particular location.
Members of the research team based in China, where primate reproductive technology is much more advanced than in the United States, injected the CRISPR components into fertilized macaque eggs, producing embryos that carried the Shank3 mutation.
Researchers at MIT, where much of the data was analyzed, found that the macaques with Shank3 mutations showed behavioral patterns similar to those seen in humans with the mutated gene. They tended to wake up frequently during the night, and they showed repetitive behaviors. They also engaged in fewer social interactions than other macaques.
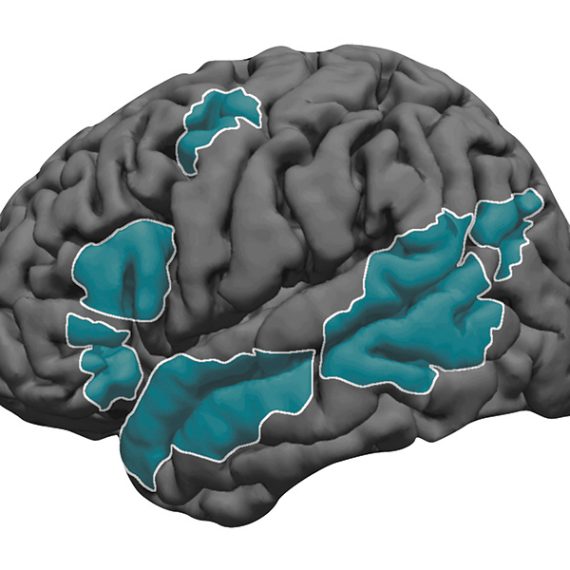
Magnetic resonance imaging (MRI) scans also revealed similar patterns to humans with autism spectrum disorder. Neurons showed reduced functional connectivity in the striatum as well as the thalamus, which relays sensory and motor signals and is also involved in sleep regulation. Meanwhile, connectivity was strengthened in other regions, including the sensory cortex.
Michael Platt, a professor of neuroscience and psychology at the University of Pennsylvania, says the macaque models should help to overcome some of the limitations of studying neurological disorders in mice, whose behavioral symptoms and underlying neurobiology are often different from those seen in humans.
“Because the macaque model shows a much more complete recapitulation of the human behavioral phenotype, I think we should stand a much greater chance of identifying the degree to which any particular therapy, whether it’s a drug or any other intervention, addresses the core symptoms,” says Platt, who was not involved in the study.
Drug development
Within the next year, the researchers hope to begin testing treatments that may affect autism-related symptoms. They also hope to identify biomarkers, such as the distinctive functional brain connectivity patterns seen in MRI scans, that would help them to evaluate whether drug treatments are having an effect.
A similar approach could also be useful for studying other types of neurological disorders caused by well-characterized genetic mutations, such as Rett Syndrome and Fragile X Syndrome. Fragile X is the most common inherited form of intellectual disability in the world, affecting about 1 in 4,000 males and 1 in 8,000 females. Rett Syndrome, which is more rare and almost exclusively affects girls, produces severe impairments in language and motor skills and can also cause seizures and breathing problems.
“Given the limitations of mouse models, patients really need this kind of advance to bring them hope,” Feng says. “We don’t know whether this will succeed in developing treatments, but we will see in the next few years how this can help us to translate some of the findings from the lab to the clinic.”
The research was funded, in part, by the Shenzhen Overseas Innovation Team Project, the Guangdong Innovative and Entrepreneurial Research Team Program, the National Key R&D Program of China, the External Cooperation Program of the Chinese Academy of Sciences, the Patrick J. McGovern Foundation, the National Natural Science Foundation of China, the Shenzhen Science, Technology Commission, the James and Patricia Poitras Center for Psychiatric Disorders Research at the McGovern Institute at MIT, the Stanley Center for Psychiatric Research at the Broad Institute of MIT and Harvard, and the Hock E. Tan and K. Lisa Yang Center for Autism Research at the McGovern Institute at MIT. The research facilities in China where the primate work was conducted are accredited by AAALAC International, a private, nonprofit organization that promotes the humane treatment of animals in science through voluntary accreditation and assessment programs.