SHERLOCK-based one-step test provides rapid and sensitive COVID-19 detection
New CRISPR-based research tool delivers results in an hour in a one-step reaction, advancing the technology closer to a point-of-care or at-home testing tool.

The team began developing tests for COVID-19 in January after learning about the emergence of a new virus which has challenged the healthcare system in China. The first version of the team’s SHERLOCK-based COVID-19 diagnostics system is already being used in hospitals in Thailand to help screen patients for COVID-19 infection.
The ability to test for COVID-19 at home, or even in pharmacies or places of employment, could be a game-changer for getting people safely back to work and into their communities.
The new test is named “STOPCovid” and is based on the STOP platform. In research it has been shown to enable rapid, accurate, and highly sensitive detection of the COVID-19 virus SARS-CoV-2 with a simple protocol that requires minimal training and uses simple, readily-available equipment, such as test tubes and water baths. STOPCovid has been validated in research settings using nasopharyngeal swabs from patients diagnosed with COVID-19. It has also been tested successfully in saliva samples to which SARS-CoV-2 RNA has been added as a proof-of-principle.
The team is posting the open protocol today on a new website, STOPCovid.science. It is being made openly available in line with the COVID-19 Technology Access Framework organized by Harvard, MIT, and Stanford. The Framework sets a model by which critically important technologies that may help prevent, diagnose, or treat COVID-19 infections may be deployed for the greatest public benefit without delay.
There is an urgent need for widespread, accurate COVID-19 testing to rapidly detect new cases, ideally without the need for specialized lab equipment. Such testing would enable early detection of new infections and drive effective “test-trace-isolate” measures to quickly contain new outbreaks. However, current testing capacity is limited by a combination of requirements for complex procedures and laboratory instrumentation and dependence on limited supplies. STOPCovid can be performed without RNA extraction, and while all patient tests have been performed with samples from nasopharyngeal swabs, preliminary experiments suggest that eventually swabs may not be necessary. Removing these barriers could help enable broad distribution.
“The ability to test for COVID-19 at home, or even in pharmacies or places of employment, could be a game-changer for getting people safely back to work and into their communities,” says Feng Zhang, a co-inventor of the CRISPR genome editing technology, an investigator at the McGovern Institute and HHMI, and a core member at the Broad Institute. “Creating a point-of-care tool is a critically important goal to allow timely decisions for protecting patients and those around them.”
To meet this need, Zhang, McGovern Fellows Omar Abudayyeh and Jonathan Gootenberg, and colleagues initiated a push to develop STOPCovid. They are sharing their findings and packaging reagents so other research teams can rapidly follow up with additional testing or development. The group is also sharing data on the StopCOVID.science website and via a submitted preprint. The website is also a hub where the public can find the latest information on the team’s developments.

Credit: Justin Knight
How it works
The STOPCovid test combines CRISPR enzymes, programmed to recognize signatures of the SARS-CoV-2 virus, with complementary amplification reagents. This combination allows detection of as few as 100 copies of SARS-CoV-2 virus in a sample. As a result, the STOPCovid test allows for rapid, accurate, and highly sensitive detection of COVID-19 that can be conducted outside clinical laboratory settings.
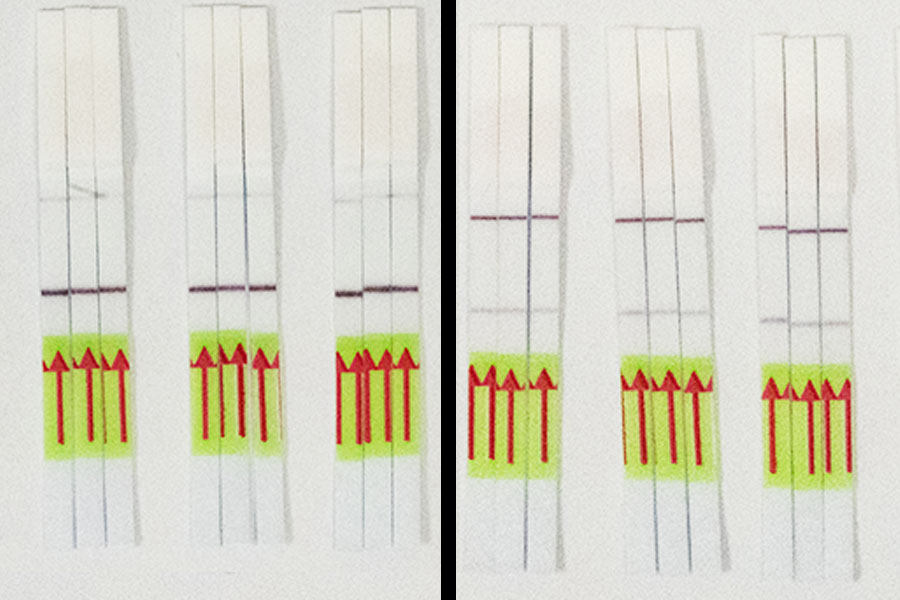
STOPCovid has been tested on patient nasopharyngeal swab in parallel with clinically-validated tests. In these head-to-head comparisons, STOPCovid detected infection with 97% sensitivity and 100% specificity. Results appear on an easy-to-read strip that is akin to a pregnancy test, in the absence of any expensive or specialized lab equipment. Moreover, the researchers spiked mock SARS-CoV-2 genomes into healthy saliva samples and showed that STOPCovid is capable of sensitive detection from saliva, which would obviate the need for swabs in short supply and potentially make sampling much easier.
“The test aims to ultimately be simple enough that anyone can operate it in low-resource settings, including in clinics, pharmacies, or workplaces, and it could potentially even be put into a turn-key format for use at home,” says Abudayyeh.
Gootenberg adds, “Since STOPCovid can work in less than an hour and does not require any specialized equipment, and if our preliminary results from testing synthetic virus in saliva bear out in patient samples, it could address the need for scalable testing to reopen our society.”

Importantly, the full test — both the viral genome amplification and subsequent detection — can be completed in a single reaction, as outlined on the website, from swabs or saliva. To engineer this, the team tested a number of CRISPR enzymes to find one that works well at the same temperature needed by the enzymes that perform the amplification. Zhang, Abudayyeh, Gootenberg and their teams, including graduate students Julia Joung and Alim Ladha, settled on a protein called AapCas12b, a CRISPR protein from the bacterium Alicyclobacillus acidophilus, responsible for the “off” taste associated with spoiled orange juice. With AapCas12b, the team was able to develop a test that can be performed at a constant temperature and does not require opening tubes midway through the process, a step that often leads to contamination and unreliable test results.
Information sharing and next steps
The team has prepared reagents for 10,000 tests to share with scientists and clinical collaborators for free around the world who want to evaluate the STOPCovid test for potential diagnostic use, and they have set up a website to share the latest data and updates with the scientific and clinical community. Kits and reagents can also be requested via a form on the website.
Acknowledgments: Patient samples were provided by Keith Jerome, Alex Greninger, Robert Bruneau, Mee-li W. Huang, Nam G. Kim, Xu Yu, Jonathan Li, and Bruce Walker. This work was supported by the Patrick J. McGovern Foundation and the McGovern Institute for Brain Research. F.Z is also supported by the NIH (1R01- MH110049 and 1DP1-HL141201 grants); Mathers Foundation; the Howard Hughes Medical Institute; Open Philanthropy Project; J. and P. Poitras; and R. Metcalfe.
Declaration of conflicts of interest: F.Z., O.O.A., J.S.G., J.J., and A.L. are inventors on patent applications related to this technology filed by the Broad Institute, with the specific aim of ensuring this technology can be made freely, widely, and rapidly available for research and deployment. O.O.A., J.S.G., and F.Z. are co-founders, scientific advisors, and hold equity interests in Sherlock Biosciences, Inc. F.Z. is also a co-founder of Editas Medicine, Beam Therapeutics, Pairwise Plants, and Arbor Biotechnologies.