New MRI probe can reveal more of the brain’s inner workings
Tracing connections between neuron populations could help researchers map brain circuits that underlie behavior and perception.
Using a novel probe for functional magnetic resonance imaging (fMRI), MIT biological engineers have devised a way to monitor individual populations of neurons and reveal how they interact with each other.
Similar to how the gears of a clock interact in specific ways to turn the clock’s hands, different parts of the brain interact to perform a variety of tasks, such as generating behavior or interpreting the world around us. The new MRI probe could potentially allow scientists to map those networks of interactions.
“With regular fMRI, we see the action of all the gears at once. But with our new technique, we can pick up individual gears that are defined by their relationship to the other gears, and that’s critical for building up a picture of the mechanism of the brain,” says Alan Jasanoff, an MIT professor of biological engineering, brain and cognitive sciences, and nuclear science and engineering.
Using this technique, which involves genetically targeting the MRI probe to specific populations of cells in animal models, the researchers were able to identify neural populations involved in a circuit that responds to rewarding stimuli. The new MRI probe could also enable studies of many other brain circuits, the researchers say.
Jasanoff, who is also an associate investigator at the McGovern Institute, is the senior author of the study, which appears today in Nature Neuroscience. The lead authors of the paper are recent MIT PhD recipient Souparno Ghosh and former MIT research scientist Nan Li.
Tracing connections
Traditional fMRI imaging measures changes to blood flow in the brain, as a proxy for neural activity. When neurons receive signals from other neurons, it triggers an influx of calcium, which causes a diffusible gas called nitric oxide to be released. Nitric oxide acts in part as a vasodilator that increases blood flow to the area.
Imaging calcium directly can offer a more precise picture of brain activity, but that type of imaging usually requires fluorescent chemicals and invasive procedures. The MIT team wanted to develop a method that could work across the brain without that type of invasiveness.
“If we want to figure out how brain-wide networks of cells and brain-wide mechanisms function, we need something that can be detected deep in tissue and preferably across the entire brain at once,” Jasanoff says. “The way that we chose to do that in this study was to essentially hijack the molecular basis of fMRI itself.”
The researchers created a genetic probe, delivered by viruses, that codes for a protein that sends out a signal whenever the neuron is active. This protein, which the researchers called NOSTIC (nitric oxide synthase for targeting image contrast), is an engineered form of an enzyme called nitric oxide synthase. The NOSTIC protein can detect elevated calcium levels that arise during neural activity; it then generates nitric oxide, leading to an artificial fMRI signal that arises only from cells that contain NOSTIC.
The probe is delivered by a virus that is injected into a particular site, after which it travels along axons of neurons that connect to that site. That way, the researchers can label every neural population that feeds into a particular location.
“When we use this virus to deliver our probe in this way, it causes the probe to be expressed in the cells that provide input to the location where we put the virus,” Jasanoff says. “Then, by performing functional imaging of those cells, we can start to measure what makes input to that region take place, or what types of input arrive at that region.”
Turning the gears
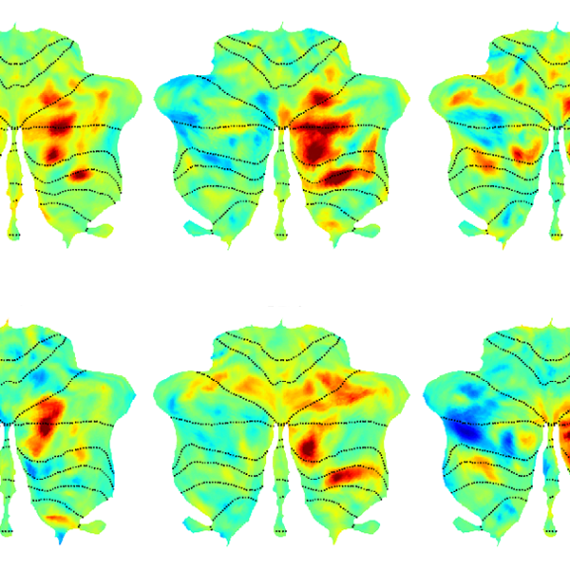
In the new study, the researchers used their probe to label populations of neurons that project to the striatum, a region that is involved in planning movement and responding to reward. In rats, they were able to determine which neural populations send input to the striatum during or immediately following a rewarding stimulus — in this case, deep brain stimulation of the lateral hypothalamus, a brain center that is involved in appetite and motivation, among other functions.
One question that researchers have had about deep brain stimulation of the lateral hypothalamus is how wide-ranging the effects are. In this study, the MIT team showed that several neural populations, located in regions including the motor cortex and the entorhinal cortex, which is involved in memory, send input into the striatum following deep brain stimulation.
“It’s not simply input from the site of the deep brain stimulation or from the cells that carry dopamine. There are these other components, both distally and locally, that shape the response, and we can put our finger on them because of the use of this probe,” Jasanoff says.
During these experiments, neurons also generate regular fMRI signals, so in order to distinguish the signals that are coming specifically from the genetically altered neurons, the researchers perform each experiment twice: once with the probe on, and once following treatment with a drug that inhibits the probe. By measuring the difference in fMRI activity between these two conditions, they can determine how much activity is present in probe-containing cells specifically.
The researchers now hope to use this approach, which they call hemogenetics, to study other networks in the brain, beginning with an effort to identify some of the regions that receive input from the striatum following deep brain stimulation.
“One of the things that’s exciting about the approach that we’re introducing is that you can imagine applying the same tool at many sites in the brain and piecing together a network of interlocking gears, which consist of these input and output relationships,” Jasanoff says. “This can lead to a broad perspective on how the brain works as an integrated whole, at the level of neural populations.”
The research was funded by the National Institutes of Health and the MIT Simons Center for the Social Brain.
Paper: "Functional dissection of neural circuitry using a genetic reporter for fMRI"