In a new brain-wide circuit tracing study, scientists at MIT’s Picower Institute for Learning and Memory focused selective attention on a circuit that governs, fittingly enough, selective attention. The comprehensive maps they produced illustrate how broadly the mammalian brain incorporates and integrates information to focus its sensory resources on its goals.
Working in mice, the team traced thousands of inputs into the circuit, a communication loop between the anterior cingulate cortex (ACC) and the lateral posterior (LP) thalamus. In primates the LP is called the pulvinar. Studies in humans and nonhuman primates have indicated that the byplay of these two regions is critical for brain functions like being able to focus on an object of interest in a crowded scene, says study co-lead author Yi Ning Leow, a graduate student in the lab of senior author Mriganka Sur, the Newton Professor in MIT’s Department of Brain and Cognitive Sciences. Research has implicated dysfunction in the circuit in attention-affecting disorders such as autism and attention deficit/hyperactivity disorder.
The new study in the Journal of Comparative Neurology extends what’s known about the circuit by detailing it in mice, Leow says, importantly showing that the mouse circuit is closely analogous to the primate version even if the LP is proportionately smaller and less evolved than the pulvinar.
“In these rodent models we were able to find very similar circuits,” Leow says. “So we can possibly study these higher-order functions in mice as well. We have a lot more genetic tools in mice so we are better able to look at this circuit.”
The study, also co-led by former MIT undergraduate Blake Zhou, therefore provides a detailed roadmap in the experimentally accessible mouse model for understanding how the ACC and LP cooperate to produce selective attention. For instance, now that Leow and Zhou have located all the inputs that are wired into the circuit, Leow is tapping into those feeds to eavesdrop on the information they are carrying. Meanwhile, she is correlating that information flow with behavior.
“This study lays the groundwork for understanding one of the most important, yet most elusive, components of brain function, namely our ability to selectively attend to one thing out of several, as well as switch attention,” Sur says.
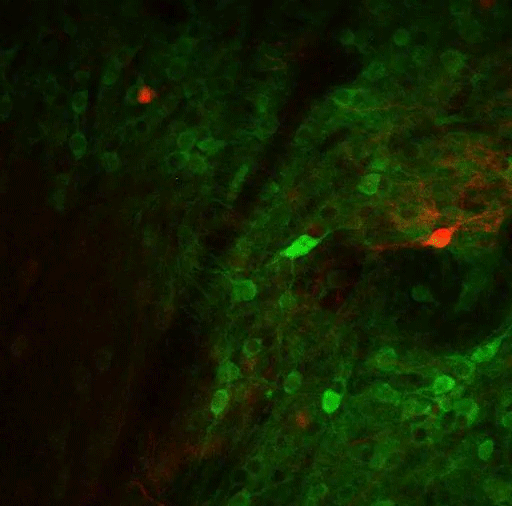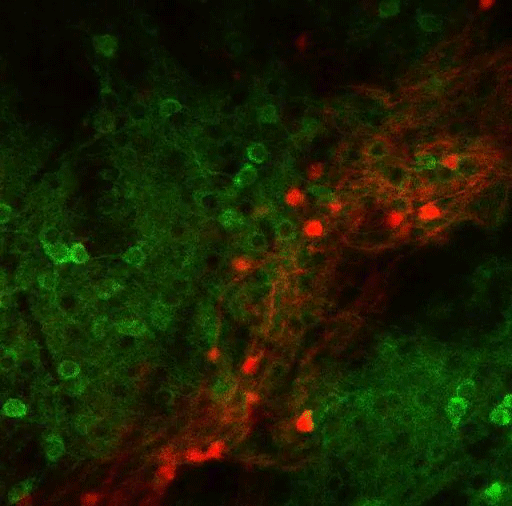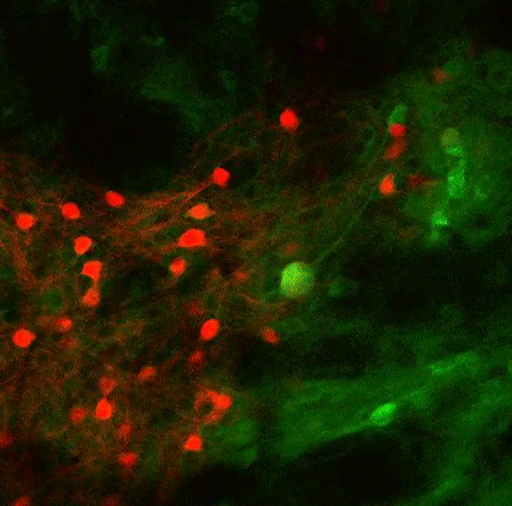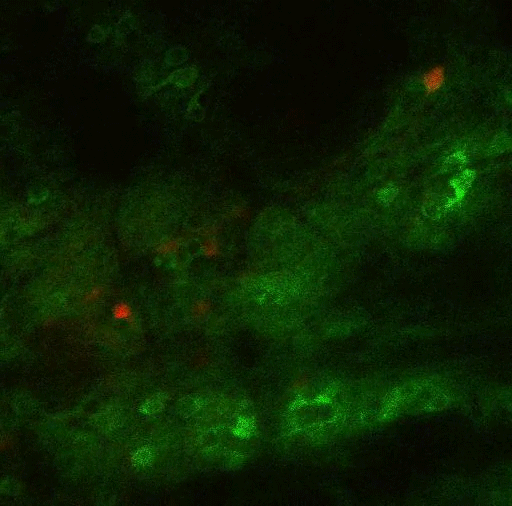
Using virally mediated circuit-tracing techniques pioneered by co-author Ian Wickersham, principal research scientist in brain and cognitive sciences and the McGovern Institute for Brain Research at MIT, the team found distinct sources of input for the ACC and the LP. Generally speaking, the detailed study finds that the majority of inputs to the ACC were from frontal cortex areas that typically govern goal-directed planning, and from higher visual areas. The bulk of inputs to the LP, meanwhile, were from deeper regions capable of providing context such as the mouse’s needs, location and spatial cues, information about movement, and general information from a mix of senses.
So even though focusing attention might seem like a matter of controlling the senses, Leow says, the circuit pulls in a lot of other information as well.
“We’re seeing that it’s not just sensory — there are so many inputs that are coming from non-sensory areas as well, both sub-cortically and cortically,” she says. “It seems to be integrating a lot of different aspects that might relate to the behavioral state of the animal at a given time. It provides a way to provide a lot of internal and special context for that sensory information.”
Given the distinct sets of inputs to each region, the ACC may be tasked with focusing attention on a desired object, while the LP is modulating how the ACC goes about making those computations, accounting for what’s going on both inside and outside the animal. Decoding just what that incoming contextual information is, and what the LP tells the ACC, are the key next steps, Leow says. Another clear set of questions the study raises are what are the circuit’s outputs. In other words, after it integrates all this information, what does it do with it?
The paper’s other authors are Heather Sullivan and Alexandria Barlowe.
A National Science Scholarship, the National Institutes of Health, and the JPB Foundation provided support for the study.