“RNA-guided biology is what lets you make programmable tools that are really easy to use. So the more we can find, the better,” says McGovern fellow Omar Abudayyeh, who led the research with McGovern fellow Jonathan Gootenberg.
CRISPR, an ancient bacterial defense system, has made it clear how useful RNA-guided enzymes can be when they are adapted for use in the lab. CRISPR-based genome editing tools developed by McGovern investigator Feng Zhang, Abudayyeh, Gootenberg and others have changed the way scientists modify DNA, accelerating research and enabling the development of many experimental gene therapies.
Researchers have since uncovered other RNA-guide enzymes throughout the bacterial world, many with features that make them valuable in the lab. The discovery of Fanzors, whose ability to cut DNA in an RNA-guided manner was reported by Zhang’s group earlier this year, opens a new frontier of RNA-guided biology. Fanzors were the first such enzymes to be found in eukaryotic organisms—a wide group of lifeforms, including plants, animals, and fungi, defined by the membrane-bound nucleus that holds each cell’s genetic material. (Bacteria, which lack nuclei, belong to a group known as prokaryotes.)
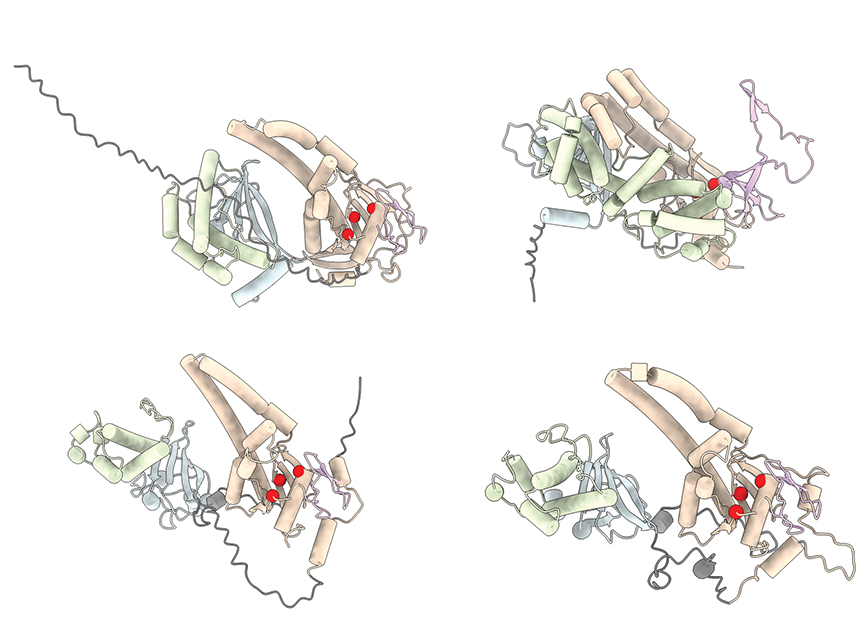
“People have been searching for interesting tools in prokaryotic systems for a long time, and I think that that has been incredibly fruitful,” says Gootenberg. “Eukaryotic systems are really just a whole new kind of playground to work in.”
One hope, Abudayyeh and Gootenberg say, is that enzymes that naturally evolved in eukaryotic organisms might be better suited to function safely and efficiently in the cells of other eukaryotic organisms, including humans. Zhang’s group has shown that Fanzor enzymes can be engineered to precisely cut specific DNA sequences in human cells. In the new work, Abudayyeh and Gootenberg discovered that some Fanzors can target DNA sequences in human cells even without optimization. “The fact that they work quite efficiently in mammalian cells was really fantastic to see,” Gootenberg says.
Prior to the current study, hundreds of Fanzors had been found among eukaryotic organisms. Through an extensive search of genetic databases led by lab member Justin Lim, Gootenberg and Abudayyeh’s team has now expanded the known diversity of these enzymes by an order of magnitude.
Among the more than 3,600 Fanzors that the team found in eukaryotes and the viruses that infect them, the researchers were able to identify five different families of the enzymes. By comparing these enzymes’ precise makeup, they found evidence of a long evolutionary history.
Fanzors likely evolved from RNA-guided DNA-cutting bacterial enzymes called TnpBs. In fact, it was Fanzors’ genetic similarities to these bacterial enzymes that first caught the attention of both Zhang’s group and Gootenberg and Abudayyeh’s team.
The evolutionary connections that Gootenberg and Abudayyeh traced suggest that these bacterial predecessors of Fanzors probably entered eukaryotic cells, initiating their evolution, more than once. Some were likely transmitted by viruses, while others may have been introduced by symbiotic bacteria. The research also suggests that after they were taken up by eukaryotes, the enzymes evolved features suited to their new environment, such as a signal that allows them to enter a cell nucleus, where they have access to DNA.
Through genetic and biochemical experiments led by graduate student Kaiyi Jiang, the team determined that Fanzors have evolved a DNA-cutting active site that is distinct from that of their bacterial predecessors. This seems to allow the enzyme to cut its target sequence more precisely the ancestors of TnpB, when targeted to a sequence of DNA in a test tube, become activated and cut other sequences in the tube; Fanzors lack this promiscuous activity. When they used an RNA guide to direct the enzymes to cut specific sites in the genome of human cells, they found that certain Fanzors were able to cut these target sequences with about 10 to 20 percent efficiency.
With further research, Abudayyeh and Gootenberg hope that a variety of sophisticated genome editing tools can be developed from Fanzors. “It’s a new platform, and they have many capabilities,” says Gootenberg. “Opening up the whole eukaryotic world to these types of RNA-guided systems is going to give us a lot to work on,” Abudayyeh adds.